Evolution of translation initiation systems in eukaryotes
In eukaryotes, translation initiation requires many factors intended to recruit the small ribosomal subunit (40S) with the tRNAMet initiator near the 5’ end of the capped mRNAs. Once the ribosome/tRNAMet/mRNA pre-initiation complex is formed, a process called scanning allows the pre-initiation complex to slide and inspect the 5’ untranslated region (UTR) of the mRNA in order to identify the initiator AUG codon. Validation of the codon-anticodon interaction by initiation factors is followed by the assembly of the large ribosomal subunit (60S) and departure of initiation factors. The assembled 80S ribosome is then ready to enter the elongation phase.
To gain in efficiency and to hijack the cellular translation components certain viral mRNA bypass the classic initiation process using “Internal Ribosome Entry Sites” (IRES) which allow them to recruit very efficiently ribosomes inside the mRNA and independently from the 5’ cap structure. IRES are also present in certain cellular mRNAs and are activated under particular cellular conditions (stress, cell cycle events).
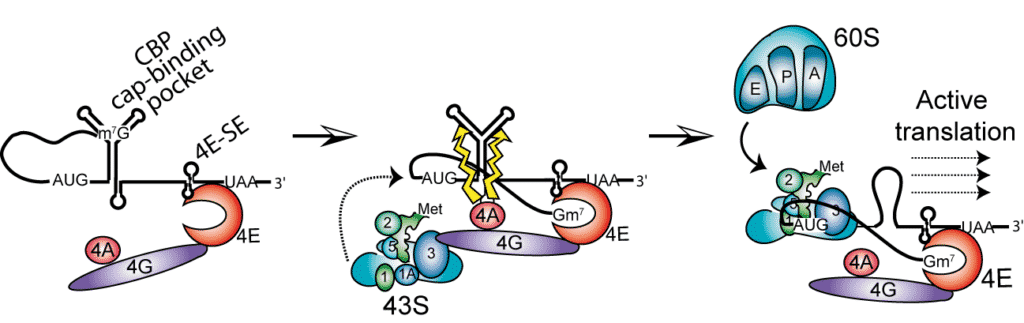
The project is focused on translation initiation of mouse histone H4 mRNA. This highly conserved mRNA exhibits very short 5’ UTR and no poly A tail. These features exclude conventional translation initiation mechanism with a “scanning” step of the UTR by the ribosome. We showed that H4 mRNA is using a new type of initiation mechanism called “tethering” allowing the mRNA of histone H4 to be very efficiently translated (Martin et al., 2011).
H4 mRNA contains in its open reading frame 2 structural elements playing critical roles during the initiation process. One element binds initiation factor eIF4E independently from the cap located on the 5’ end. From this site, the activated 43S small ribosomal subunit is recruited by eIF4F and directly loaded on the AUG initiation codon without any scanning of the mRNA by the ribosome. The second essential structure of the mRNA is located 19 nucleotides downstream the AUG codon. This element is folded as a three-way helix junction domain and facilitates the positioning of the ribosome on the AUG codon. Then, the 60S is assembled, forming the 80S particle ready to start translation. This basic mechanism of “tethering” ensures an immediate start of the translation without the slow and cost-effective step of scanning (Martin et al., 2011).
See more: « Quand la cellule se prend pour un virus »

 Intranet
Intranet Access
Access Contact
Contact